Core Workflow
iMano's Core Workflow
Pipeline Overview
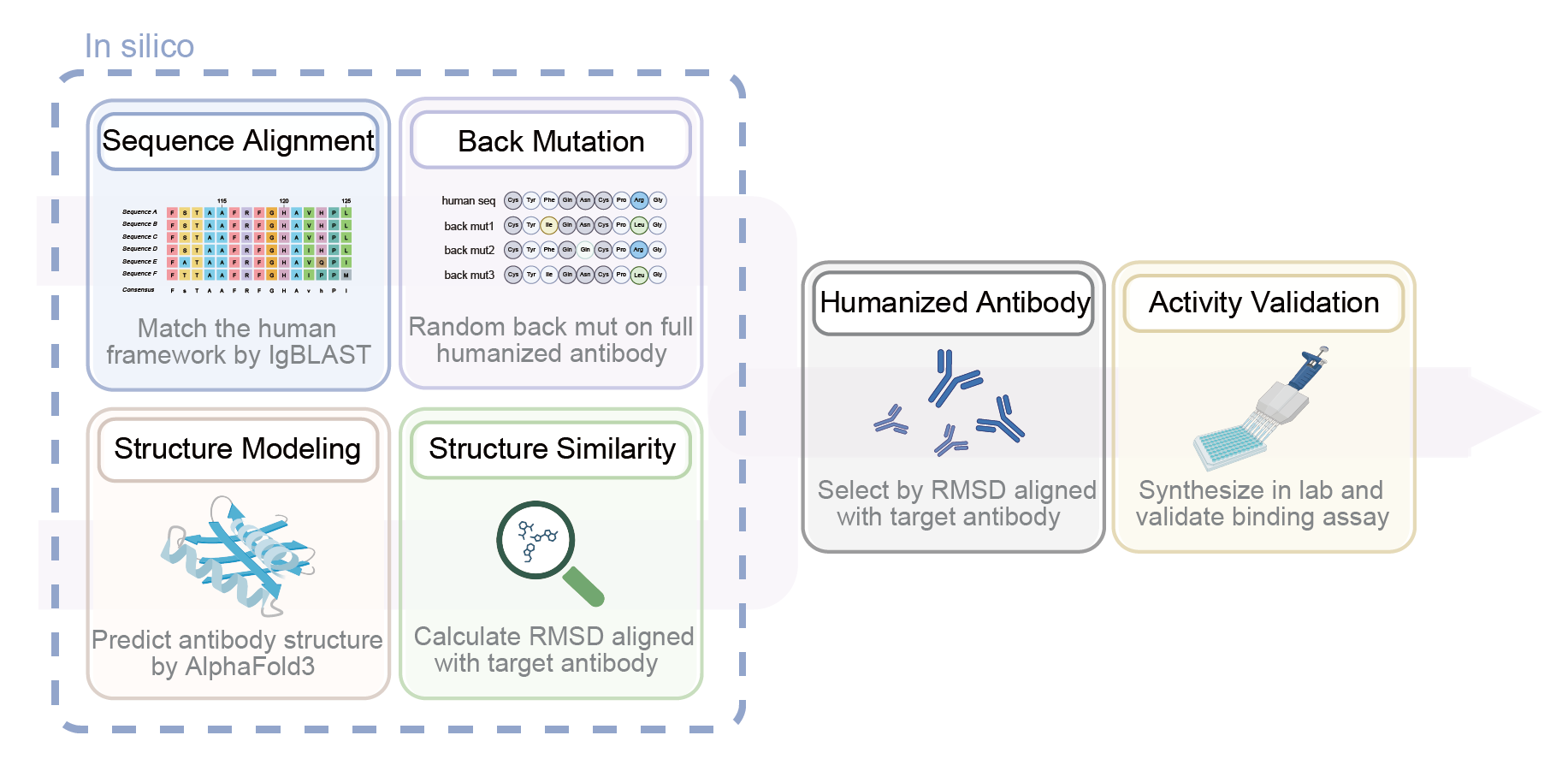
Key steps: ① IgBLAST germline matching → ② AlphaFold3 structure prediction → ③ RMSD screening → ④ Binding validation.
Residue Selection & Candidate Generation
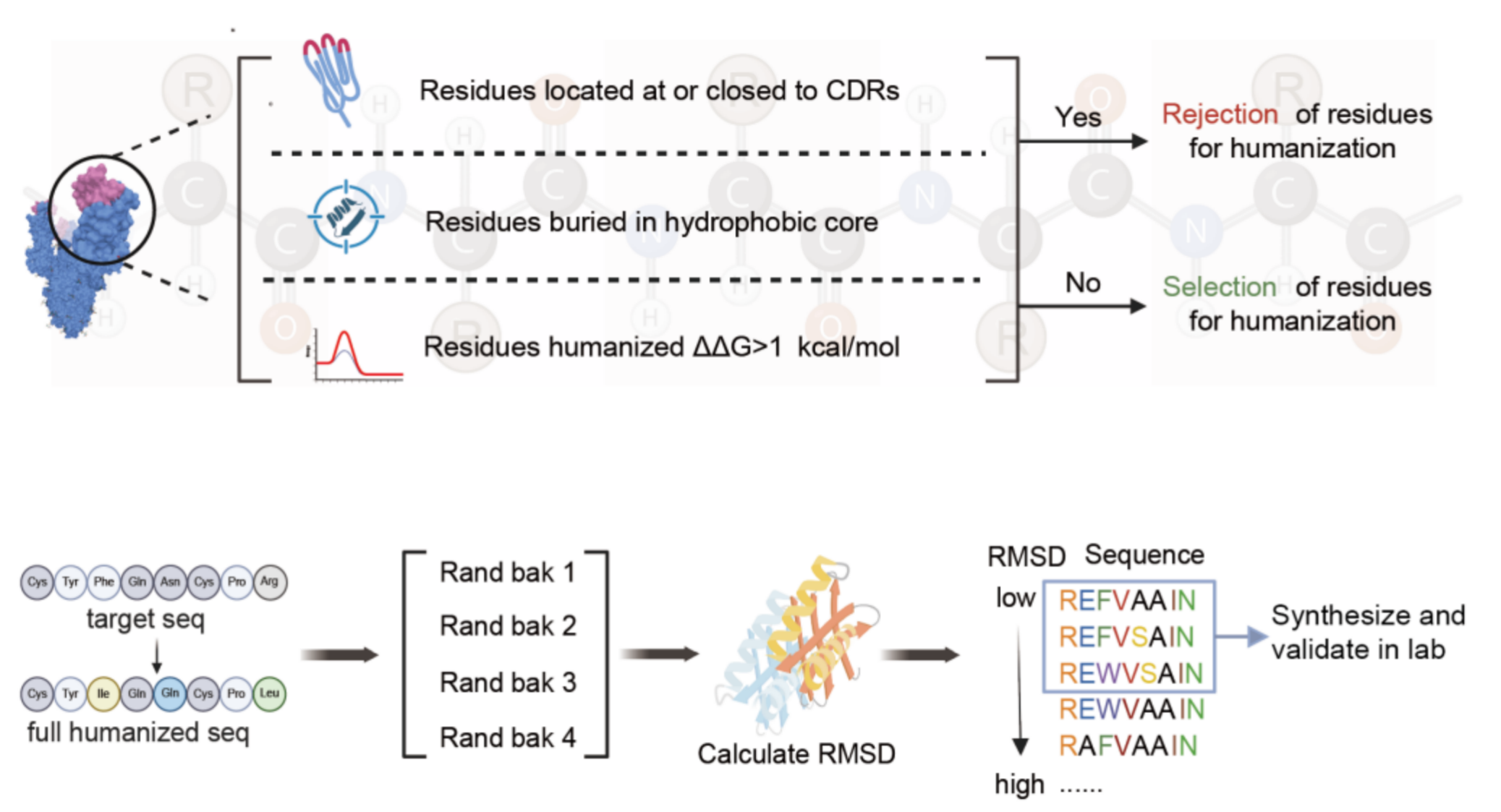
Core logic: Fix hydrophobic core residues (ΔΔG < 1 kcal/mol) + max 4 back mutations per chain + prioritize low CDR RMSD candidates.