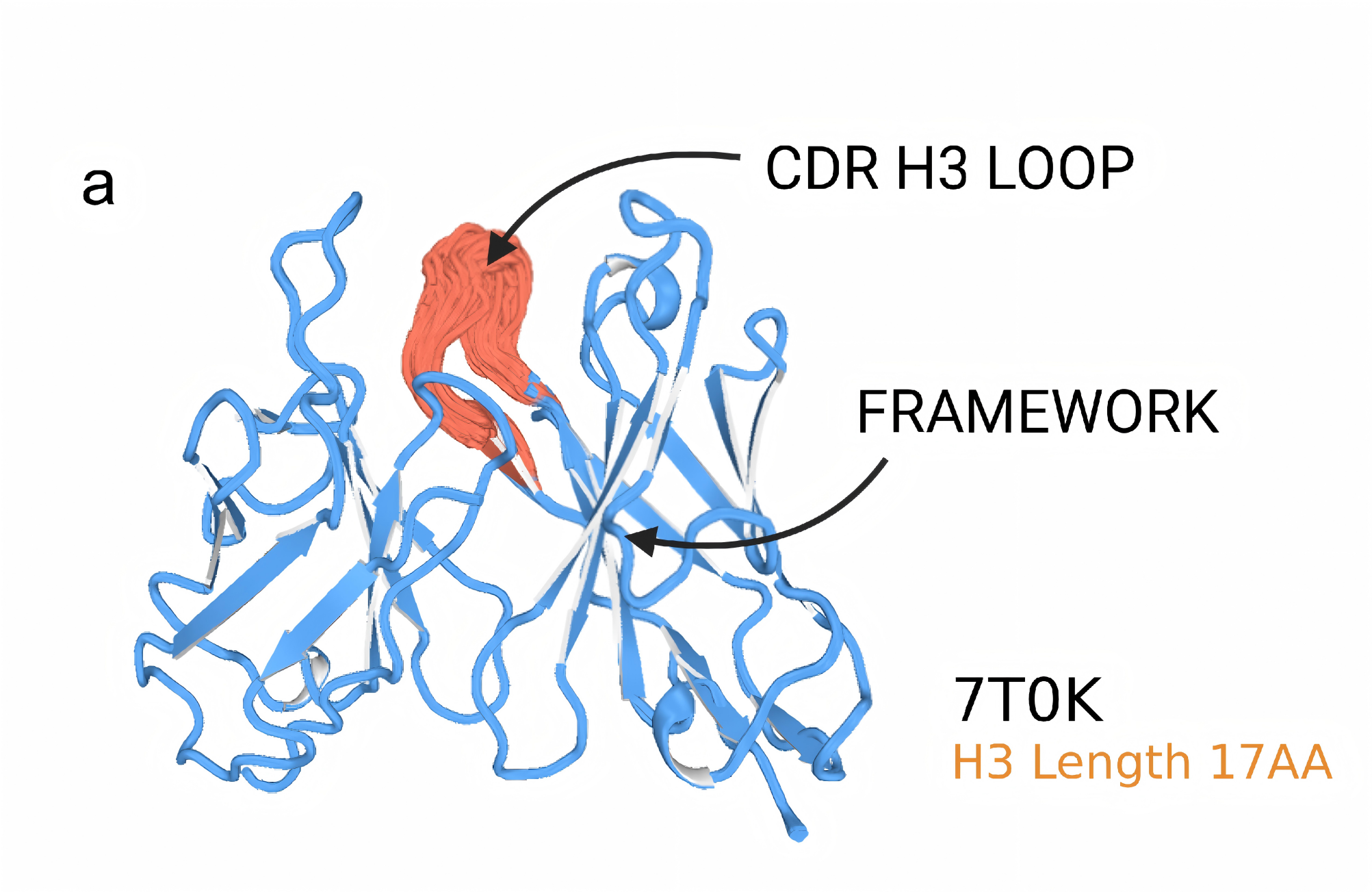
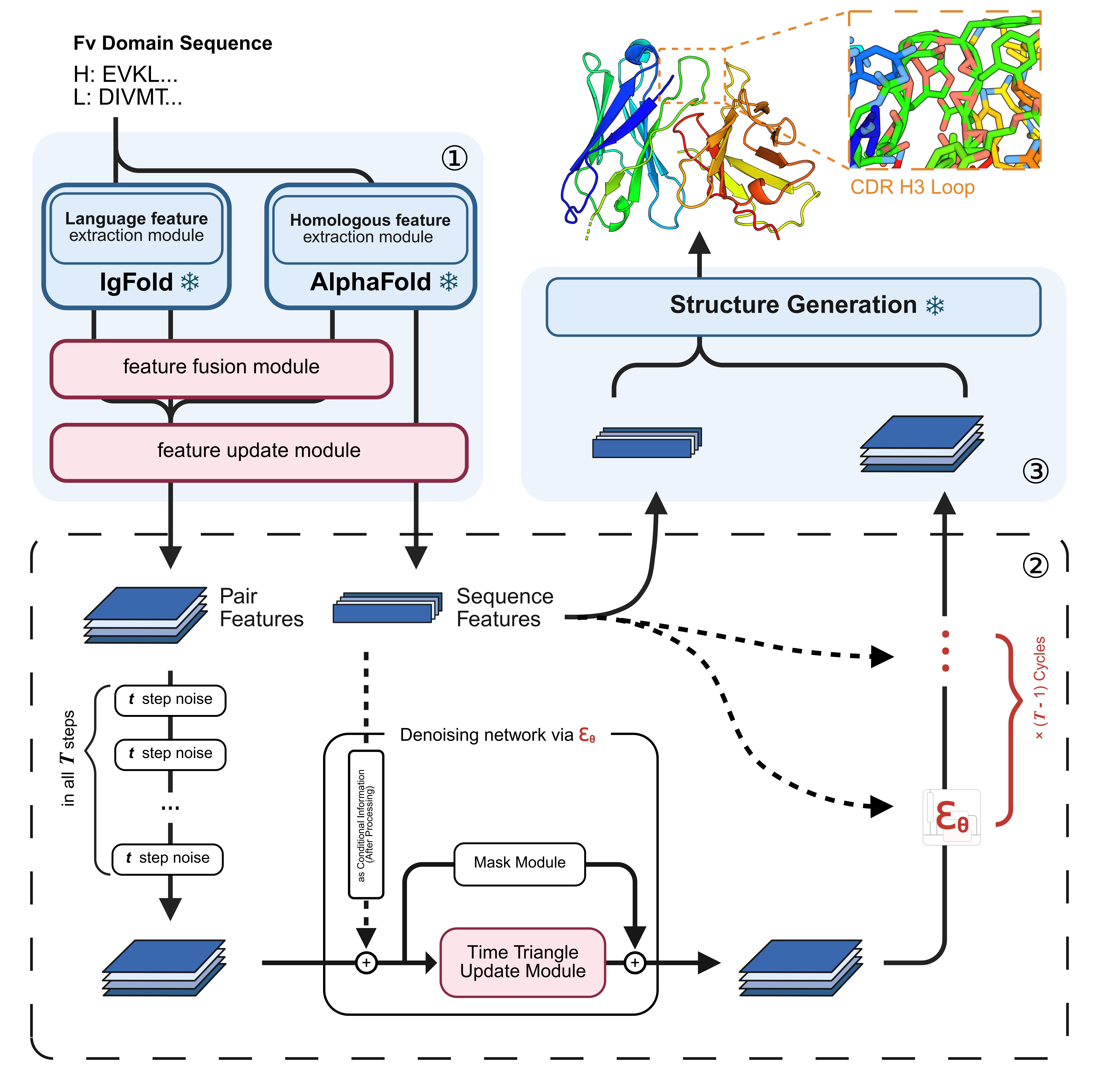
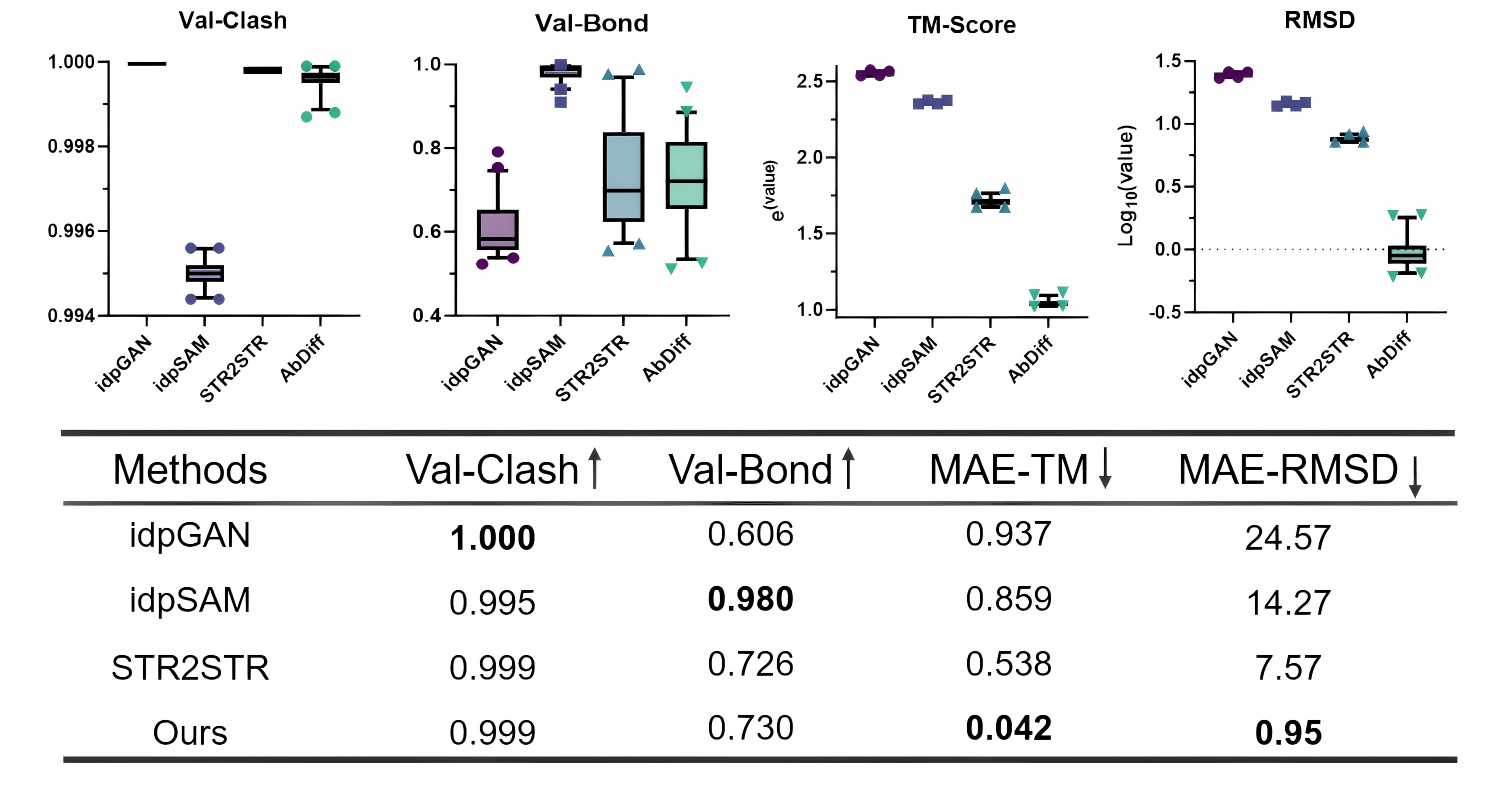
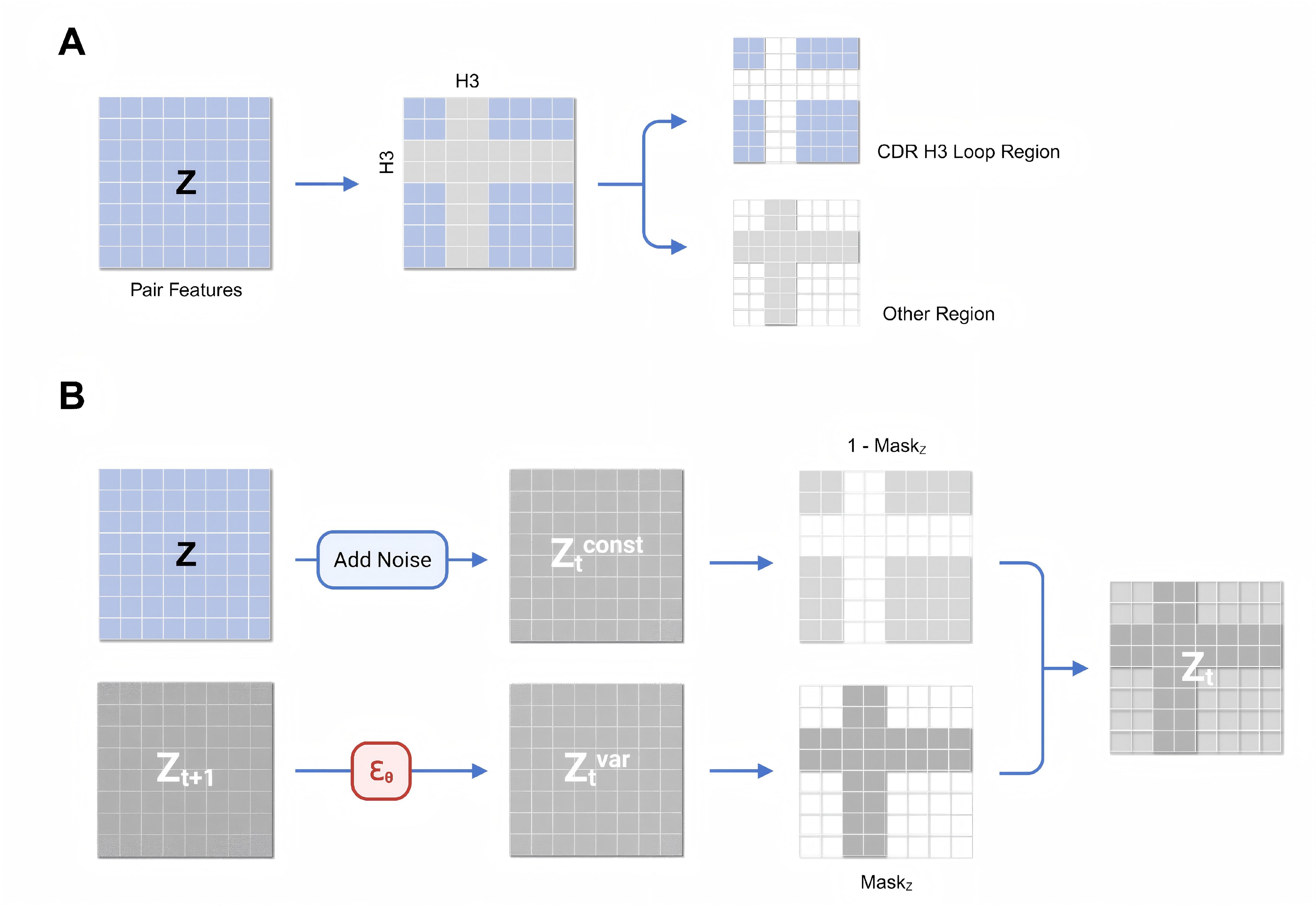
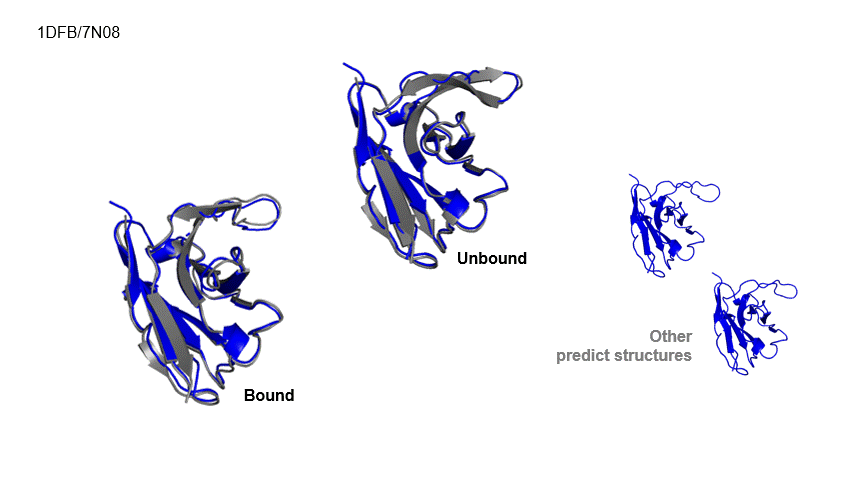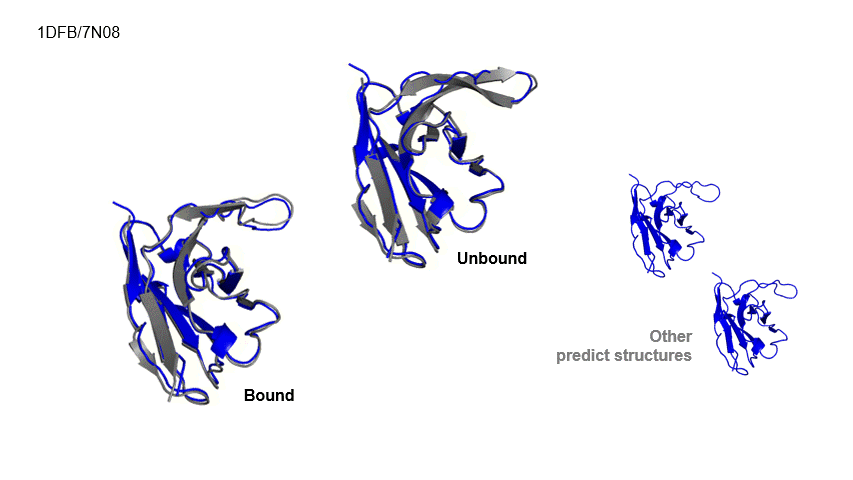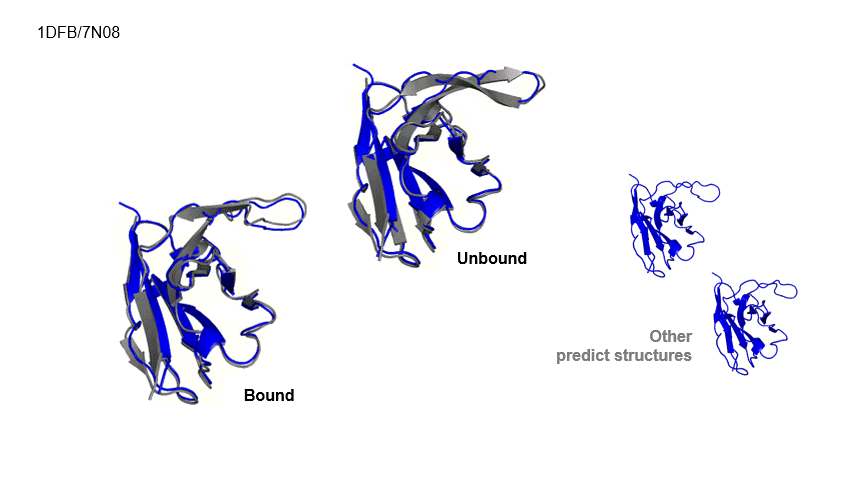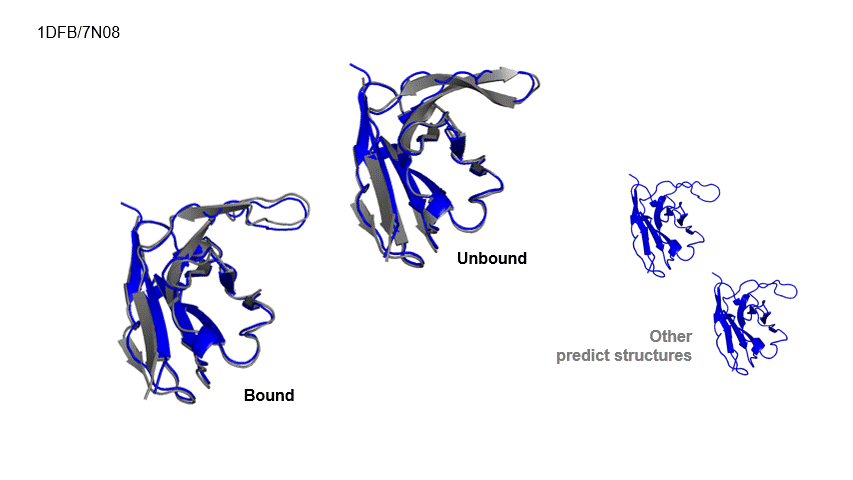
Generative Prediction of Antibody Conformational Ensembles Using Denoising Diffusion Probabilistic Models
This work is being submitted.
Chao Peng1,#,
Ziming Suo2,#,
Zelong Wang1,
Ali Nobakhtnamini1,
Yi Feng2,
Juncheng Qian2,
Yu Kong2,
Chuang Qin2,
Zifei Wang2,
Yanling Wu2,3,*,
Weifeng Ge1,*,
Tianlei Ying2,3,4,*
1 College of Computer Science and Artificial Intelligence, Fudan University, Shanghai, China. 2 Key Laboratory of Medical Molecular Virology (MOE/NHC/CAMS), Shanghai Institute of Infectious Disease and Biosecurity, School of Basic Medical Sciences, Department of Pulmonary and Critical Care Medicine, Department of Liver Surgery and Transplantation, Zhongshan Hospital, Fudan University, Shanghai, China. 3 Shanghai Engineering Research Center for Synthetic Immunology and Shanghai Key Laboratory of Lung Inflammation and Injury, Shanghai, China. 4 Shanghai Innovation Institute, Shanghai, China. # These authors contributed equally. * Corresponding author.